Docket #: S11-282
Microfluidic Cell Sorting and Bacterial Detection with Polymeric Film Imprinting
Researchers in Prof. Dick Zare's laboratory have developed an efficient, label-free platform to separate and detect cells, bacteria, or other particles in fluid samples. This method employs cell/particle cavities created in a polymeric film (imprinted polymeric film, IPF). When incorporated into microfluidic chips, the IPFs chemically recognize particles or cells of interest in a sample and selectively re-adsorb them within minutes. Unlike FACS (fluorescence-activated cell sorting), the IPF-based technology can sort multiple cells simultaneously and does not require fluorescent labels. This method is well-suited for recognizing bacterial cells or other particles that are too large for chromatography. This technique could be applied to cell sorting from any fluid sample (such as water, blood, urine or plasma) for research or diagnostics.
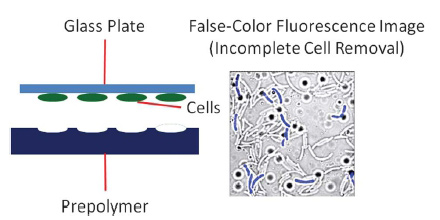
Schematic (left) and confocal microscopy image (right) of cell imprints on a polymer surface. The IPF is generated by pressing bacteria adhered on a glass plate into a prepolymer. After the polymer is cured and the glass plate removed. The remaining cavities resemble the cells in size and shape. (Remaining cells can be completely removed by sonication.)
Stage of Research
The inventors have demonstrated this technology using IPF templates in a microfluidic chip to separate strains of cyanobacteria with 80-90% efficiency. The inventors continue to research the platform as a diagnostic tool for tuberculosis and are developing modifications to improve safety and optimize detection.
Applications
- Diagnostics - identify bacteria or other pathogens from bodily fluids (e.g. blood, urine, sputum)
- Cell sorting - isolate cells with unique characteristics (e.g. infected blood or rare metastasizing tumor cells) for characterizing disease states
Advantages
- Label-free - cell extraction does not require size differential, fluorescence, magnetic label or any other modification of the cells being analyzed
- High-throughput - multiple cells can be sorted simultaneously (unlike FACS, where sorting occurs cell-by-cell) within minutes
- Detection in unique size range - can identify cells (or possibly other bio analytes) that are too large for chromatography
- Fast and low cost - microfluidic-based analysis would be faster and less expensive than culturing bacteria or microscopy
- Specific and efficient - initial proof-of-principle demonstrated separation of different strains of cyanobacteria with 80-90% efficiency, despite a minimal difference in morphology
Publications
- Kangning Ren and Richard N. Zare, "Chemical Recognition in Cell-2 Imprinted Polymers", ACS Nano, published online April 2nd, 2012.
- Romana Schirhagl, Eric W. Hall, Ingo Fuereder and Richard N. Zare, "Separation of bacteria with imprinted polymeric films", Analyst, 2012,137, pp. 1495-1499, DOI: 10.1039/c2an15927
- Romana Schirhagl and Richard Zare, "Surface-Imprinted Polymers in Microfluidic Devices", Science China Chemistry Apr. 2012, Vol. 55, (4) 469-483.
Related Links
Patents
- Published Application: 20130309657
- Issued: 9,557,250 (USA)
Similar Technologies
-
Microfluidics device for high-throughput liquid biopsy and other cellular analyses S17-169Microfluidics device for high-throughput liquid biopsy and other cellular analyses
-
Trapping Nanoscale Objects in Solution S04-213Trapping Nanoscale Objects in Solution
-
Punchcard Programmable Microfluidics S12-267Punchcard Programmable Microfluidics