Docket #: S24-151
Droplet Arrays For High-Throughput Protein Analysis and Screening
Stanford researchers have developed a novel methodology for the high-throughput expression and kinetic characterization of numerous enzyme variants in parallel using microfluidic droplet arrays.
Enzymes play a critical role in medicine and industry as they can facilitate essential biochemical reactions for diagnostic assays, drug development, sustainable practices in biocatalysis and food production. However, the rapid screening of enzyme variants—an important aspect of enhancing enzyme functionality and advancing biotechnological applications—has been hindered by current methods that often struggle with scalability and require specialized equipment and costly materials.
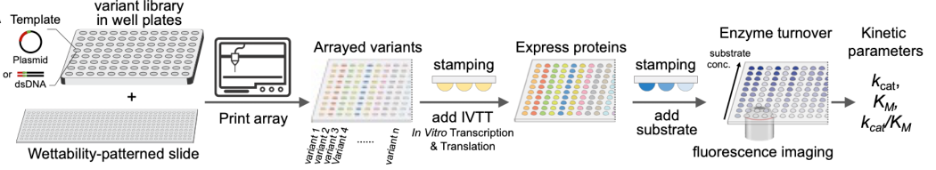
The workflow begins by printing enzyme variant libraries onto patterned hydrophilic spots, followed by stamping with protein expression reagents to produce proteins in each droplet. Substrates are then added to initiate parallel enzymatic reactions, allowing for enzyme kinetic analysis for all variants in parallel. This is a valuable tool for rapid screening and optimization of enzymes, which could accelerate product development.
Figure
Stage of Development
Proof of concept – in vitro data
Applications
- High-throughput screening of large enzyme libraries in one assay
- Analysis of enzyme activity in parallel
- Efficient drug development and diagnostics
Advantages
- Simultaneous high-throughput screening of over 1,100 enzyme variants
- Minimal reagent use, lowering costs and waste.
- 250x cheaper than robotic systems, with simple setup requiring no specialized training
Publications
- Byungjin Lee, Fanny Sunden, Michael Miller, Bumshik Pak, Anker Krebber, Stefan Lutz, Polly Morrell Fordyce. Hydrophilic/ Omniphobic droplet arrays for high-throughput and quantitative enzymology. bioRxiv 2024.07.19.604368.
Related Links
Patents
- Published Application: WO2026019982
Similar Technologies
-
Rapid Microbe-independent Plasmid Cloning and Analysis in Mammalian Cells S26-035Rapid Microbe-independent Plasmid Cloning and Analysis in Mammalian Cells
-
Paperfuge: a low-cost, high-speed, human-powered centrifuge S15-444Paperfuge: a low-cost, high-speed, human-powered centrifuge
-
Trapping Nanoscale Objects in Solution S04-213Trapping Nanoscale Objects in Solution